Our bodies rely on DNA to function, it’s often described as “the secret of life”. A computer program that describes how to make a man. However inaccurate these analogies might be, DNA is fundamental to life. In order for organisms to grown and replicate they therefore need to copy their DNA.
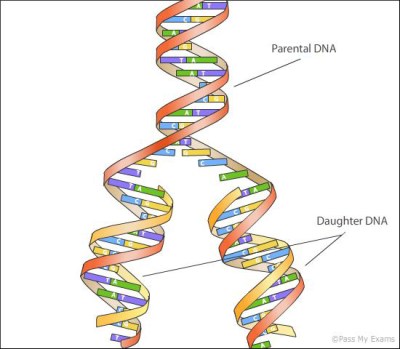
Since the discovery of its structure in 1953, the approximate method used to copy DNA has been obvious. The information in DNA is encoded in 4 nucleotides (which in their short form we call A,T,G, and C). These couple with each other in pairs, forming 2 complimentary strands that mirror each other. This structure naturally lends itself to replication. The two strands can dissociate (under heat we call this melting), and new strands form around each single stranded template.
However, this replication process can’t happen all by itself, it requires assistance. And it wasn’t until we discovered an enzyme called the DNA polymerase that we understood how this worked. In conjunction with other enzymes, double stranded DNA is unwound into 2 single strands which are replicated by the polymerase.
It’s a beautiful process (and I highly recommend Eric Lander’s lecture on the subject).
Outside of the body, we often want to amplify DNA for other purposes. That is, take a small amount of DNA and copy it so we have a large number of identical copies for further experimentation. In 1983 Kary Mullis invented a process to accomplish exactly this, for which he won the Nobel Prize, crediting his success to the use of LSD:
The Polymerase Chain Reaction (PCR) copies a single strand of DNA by first attaching primers. These are short fragments of synthetic DNA, designed to be complimentary to the original strand. The polymerase can’t work without a small fragment to start it off. With its work begun the polymerase finds it easier to continue to replicate the strand when supplied with free single nucleotides to add to the strand. With this in place the polymerase races along adding 1000s of bases per second to the original strand. In this way, one strand of DNA is made into two.
Thermal Cycling
The key to using this copying method to create large amounts of DNA is called thermal cycling. By heating the DNA melts it (breaks the double stranded DNA into 2 single strands). The heat unfortunately kills most polymerases so we need to reduce the temperature after melting and add some more (there’s a trick to avoid this that we’ll talk about later). After adding additional polymerase, primers again attach to these new single strands, and the process begins to again exponentially copy the fragments. This heating and cooling process is preformed cyclically, and the number of fragments of DNA increases exponentially. If there was 1 fragment at first there will be 2,4,8,16,32,64,128 fragments as the cycles progress.
I mentioned that the thermal cycling process kills the polymerase. This was the case with the first known polymerase, polymerase I. This is hugely annoying, as without this the process could be trivially automated and cycles could proceed just by cycling the heat. So scientists went searching for polymerases that could survive the high temperatures involved in melting DNA. Logically enough they began with the extremeophiles, creatures that love nature’s most extreme environments. The first place they looked was natural hot springs, this resulted in the discovery of a polymerase in a bacteria called Thermus aquaticus, and they named the polymerase Taq. Taq has become a standard polymerase for use in PCR, and has made scientists pretty happy:
But always unsatisfied scientists have searched out even more robust polymerases, extracting them from deep sea hydrothermal vents. Polymereases have been discovered and engineered to work with weird and wonderful artificial bases, and have enabled advances in the incorporation of labelled DNA, forming the basis of modern DNA sequencing (reading) techniques.
The Essential Equipment
 With these new polymerases in place, the PCR process becomes a trivial matter of cycling the temperature with the correct reagents in place. Commercial thermal cyclers are a strandard piece of lab equipment. But hackers have developed a number of open source thermal cyclers too, such as the openPCR or even cheaper hacked together alternatives.
With these new polymerases in place, the PCR process becomes a trivial matter of cycling the temperature with the correct reagents in place. Commercial thermal cyclers are a strandard piece of lab equipment. But hackers have developed a number of open source thermal cyclers too, such as the openPCR or even cheaper hacked together alternatives.
While PCR is a great technique for amplifying DNA there are also other useful things you can do with it too. By selecting primers carefully you can amplify a specific region in your sample. This is the basis of a technique called DNA fingerprinting.
In DNA fingerprinting PCR is used to amplify 13 specific regions in your genome. These regions are composed of repeated sequences that vary in length. Because the lengths of these regions vary from person to person, they form a unique marker.
With a sample which now contains copies of these 13 amplified regions you can now use a process called gel electrophoresis to determine their length and compare it to a known fingerprint. A while back OpenPCR described how this technique might have been used to identify Osama bin Laden.
Not only is the polymerase fundamental to life, through PCR it has become an indispensable tool for molecular biologists. As biohacking has emerged, tools like OpenPCR have opened up these techniques to a wider audience. As the biohacking community grows, hopefully we’ll see the polymerase hacked in more surprising and interesting ways.















I’d say the greatest room for innovation is lowering the barrier to entry in designing real-time pcr assays. If you could make an arduino like IDE for PCR primer design and probe design, you would start to see PCR test libraries form like you would open source software.
Outside of the body, we often what to amplify DNA for other purposes. Want*?
Thanks, fixed.
Can’t wait for the day when anybody can start hacking the human germline in the comfort of their basement lab
Ever since the day of the radioactive boy-scout, we have been at that day.
I was designing an open PCR machine like this before. The biggest limitation in the DIY designs that you see is 1) limited sample capacity (only 1-2 tubes in most of these) and ramping speed to get to the desired temperature. In practice there is a big difference in usability between something that spends 1 hour to hit all your cycles or 3 hours, after you factor the time it takes to hit each temperature. I was using a peltier like commercial units do and built a mini aluminum forge to cast the sample holder, I should pick up where I left off on that project.
Easiest way would be based on the oldest, keep reservoirs of water at each temperature and pump through the tube holder in sequence (rather than dipping in a water bath at each temp). Should be able to get cycling real fast that way. Also you don’t waste as much power pumping heat in and out of the tube holder.
Why not try resistive heating?
Resistive heating is easier and I plan on using it for the heated lid, but I wanted to be able to ramp temperature down faster than just a fan and a heatsink so I am employing a current switching design to make the peltier cool rapidly. The temperature transition has been really quick on the bare peltier, I just have to see how fast it can heat/cool the block.
Yeah, That seems reasonable. I would just see how much mass you could remove from the block and keep the control loop stable.
If I would make a PCR thermocycler, I certainly would make a block for 96-well plates or at least 8-tubes-strips. I would also add a heated lid to avoid condensation. I think peltier elements are fine.
But, really, I do not have any application for a PCR cycler at home.
Left-handed DNA .. One of my peeves.
https://hackaday.com/wp-content/uploads/2016/03/pcr.jpg?w=1824
Could one of the editors please replace thst graphic with a mirror image?
By the way, the thermosiphon PCR design seems tough to beat for speed, simplicity, cost, and scalability. Want more samples?/ Add more.tubes.
https://hackaday.io/project/1864-5-dna-replicator
This is awesome. I had no idea and find it super interesting (first google hit on “dna only twists one direction”
This will be fixed shortly.
That looks much better .. Always a nice feeling to cure neurotic impulses w/o medication. Ha!
DNA does not /only/ twist one way, left-handed DNA is a real thing, and hotly debated by some as to how prevalent it is in various natural states (especially of interest is in-vivo/in-situ conformation, the understanding of which is being greatly advanced by cryoelectrotomography)
See this hall-of-fame:
http://users.fred.net/tds/leftdna/LeftHanded.DNA.html
This is even more fuel for the fire (though I believe the author of this site is simply outdated, as I’ve gained through conversation with him in the past year)
http://notahelix.net/
Interesting ideas, but they ignore a great deal of experimental evidence regarding how DNA works in the biological context.
Unfortunately hackaday either is holding my previous (related) comments in some spam-prevention queue, or the site lost my comments :(
As I mentioned, in my talks with the author, he seems to have been too far from academia and didn’t hear about more recent techniques for in-situ/in-vivo investigation. Late-gen tech has been using cryoelectrotomography to look at DNA in its native state (with increasing resolution every year).
> Unfortunately hackaday either is holding my previous (related) comments in some spam-prevention queue, or the site lost my comments :(
spam queue. They’re out now, but I can’t figure out how they got there.
Nathan: Thanks for the clarification. I now see I misread part of your comment to which I had replied.
Well looks like my first comment got lost… but left-handed DNA is a real thing, which is being more and more intensely studied because of arguments over how DNA looks and acts in-vivo/in-situ.
http://users.fred.net/tds/leftdna/LeftHanded.DNA.html
Luckily we’ve been making great advances thanks in great part to cryoelectrotomography, and if you look that up, then also take a look at serial block-face scanning electron microscopy. It turns out we still are in the middle/late dark-ages, but things are really starting to take off! I expect this intense wave will still be picking up speed for another 50-75 years, and not crest until after then.
yes, but Z-DNA is different from every day right-handed DNA .. while is twists in a left-handed manner, its overall confirmation is also fairly different — i.e. it is not “the same as right-handed DNA, only the other direction”.
And, yeah, there is considerable work on taking the basic language of nucleotides and radically extending it.
That… That’s because we’re developing reverse chirality yeast. Have a bucket of it over here. Combine that with chemical production of sugar, and you have l-sucrose. Tastes the same, but your body can’t process it. The ultimate sugar replacement. Trillion dollar idea.
Yeah, that’s the reason the art is wrong.
Careful there. Simulating taste buds primes your digestive response and then you withhold the actual sugar. As a metabolism biologist I assure you this is a terrible idea, I’m still amazed how many people think no-calorie sweeteners are better for you then the calorie version gram for gram.
Not to forget the osmotic pressure.
And adding indigestible sugars to your gut makes for… well, you know what happens with sugar-free gummibears.
Can’t use it as rocket fuel?
I’m intrigued and this is a brilliant idea. I do PCR all the time though and volumes are typically in the 25uL range. How do you siphon that much volume around efficiently without losing a lot to the tubing? Do you just scale up the working volume and add more enzyme? Electricity cost is way less than more enzyme, especially if you use use the PCR machine for applications other than basic DNA amplification.
I wonder if the project-creator knows that a (biohacker-friendly) startup surrounding this kind of tech already tried and failed:
https://lavaamp.wordpress.com/
and lots of discussion on background, pros, and ultimately the cons that detract from practical hobby or commercial use:
Electronic requirements for redesign of Arduino PCR thermal cycler
https://groups.google.com/d/msg/diybio/x0Rg4iamuzU/HvsUlVQYlbUJ
What’s wrong with open source PCRs
https://groups.google.com/d/msg/diybio/euT3tm4fA_g/iLRTin5vBgAJ
(for those that don’t want to read forum posts:
https://www.researchgate.net/profile/Nitin_Agrawal6/publication/6365283_A_pocket-sized_convective_PCR_thermocycler/links/0912f50646768af845000000.pdf
)
Very cool to read an article largely about Rob Carlson .. he’s an interesting guy.
Prices are pretty much headed in one direction, and more and more folks will be able to dabble in bio if they want to. BentoBio just launched their kick-starter — basically a thermo-cycler, centrifuge and gel-electrophoresis kit in one small package. At $800,they’re not giving it away, but it’s cheap in comparison to the status quo.
Project Creator here!
I found about this project after I had started mine, so luckily I didn’t know about their failure or I wouldn’t have tried!
My PCR device does actually work, and the prototype has amplified up to a 1.5kb fragment, although I just haven’t tried longer fragments, so it is possible that it could also manage that. For smaller stuff, say 100-800bp, it works really fast, with about a 30-60 minute reaction generating really good bands on a gel.
When I have time again (when the new baby is sleeping through the nite), I’ll finish up the project :)
Can we maybe get some articles about making yogurt or other trivial matters? What qualifies this stuff for had? “Biohacking”? Wasn’t that just biology as done by ignorant people?
Yeah, I’d be down with some yogurt making articles. Creating food with living organisms… we do have a food hacking category.
(does a quick search)
DIY Incubator Cooks Bacteria… Or Yogurt!
Learn bio or get left behind .. Computers were once confined to labs, so was.networking .. look what happened once things opened up.
Something similar is happening to bio .. Cheap DNA, cheap computing power, cheap digital fabrication, easy access to research, and cheap sensors and actuators etc. are making it increasingly easy to do interesting things using biology as an part of the hack.
Don’t know, i found biohacking quite scary. Can’t somebody use this one day to make some super-virus or stuff like that?
You can do the the traditional way of selective breeding with even less…it’s just slower…
Except it isn’t even really all that much slower… actually might be faster, since we don’t yet know what it takes to make a virus fatal to humans in the general sense.
https://www.rt.com/news/bird-flu-killer-strain-119/
Well, that was kinda prophetic
Back in 1976 I took a 2 week NBC course (Nuclear, Biological, Chemical) with the U.S. Army.
The instructor told us that any nation with a brewery can develop biological weapons.
Took one in ’04; they’re saying it’s even easier now without the brewery. Sad part is, doing much of the number-crunching, the fan out rate has increased due to changes in climate/weather. Of course, this has also increased the ease of use and increased growth rates. Scary stuff.
Sure, somebody could make super viri or bacteria, but we can also make improvements to the immune system and such. As for “super-humans”, if everyone has the “powers”, then it doesn’t harm anyone any more than the current situation.
This article is wrong. Thermus aquaticus was not discovered when scientists started looking in hot springs because they needed enzymes, T. aquaticus was discovered in a study of the ecology of photosynthetic bacteria in Yellowstone’s hot springs.
The discovery of T. aquaticus is one of the best examples of the value of basic research:
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1208068/pdf/ge14641207.pdf
Thanks for the link, it was an interesting read!
So, how long before this happens?
1) Sneak a DNA sample from a mark [maybe the local chief of police].
2) Amplify until you have copious quantities of the marks DNA.
3) Commit crime.
4) Liberally spray the crime scene with marks DNA.
5) Profit!
It would be easier and cheaper just to liberally spray peroxide or bleach all over the crime scene. Otherwise they Could find your DNA and the police chief’s. Peroxide is preferred, bleach has an obvious odor.
Ahemmm… https://hackaday.io/project/1864-5-dna-replicator