While DNA-based computing may not be taking over silicon quite so soon, there is progress in the works. In a paper published by Small, researchers from the University of Rochester demonstrate a molecular computing system capable of calculating square roots of integers up to 900. The computer is built from synthetic biochemical logic gates using hybridization, a process where two strands of DNA join to form double-stranded DNA, and strand displacement reactions.
DNA-based circuits have already been shown to implement complex logic functions, but most existing circuits prior to the recent paper were unable to calculate square root operations. This required 4-bit binary numbers – the new prototype implements a 10-bit square root logic circuit, operating up to the decimal integer 900.
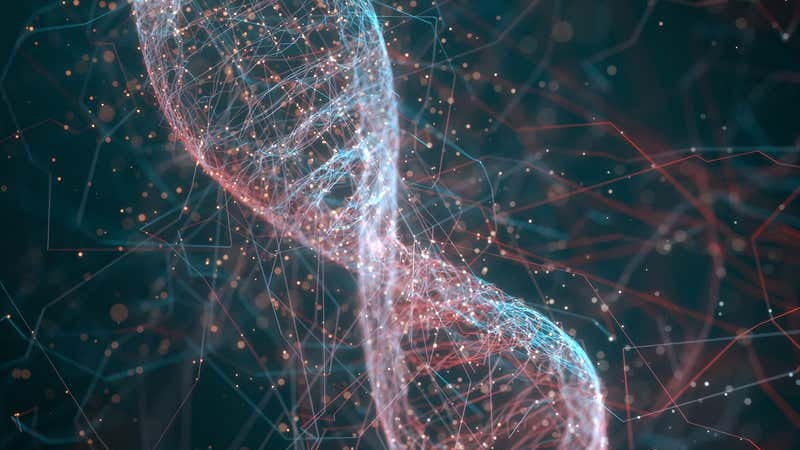
The computer uses 32 strands of DNA for storing and processing information. The process uses three modules, starting off with encoding a number on the DNA. Each combination is attached to a florescent marker, which changes signal during hybridization in the second module. The process for calculating the square root controls the signals, with the results deducted from the final color according to a threshold set in the third module.
We’re beginning to see the end of Moore’s Law approaching, with companies like Intel and AMD struggling to shrink transistors 10 nm wide. Nevertheless, with DNA molecules still about 10 time smaller than the best transistors today and DNA computing systems continuing to gain in sophistication, biochemical circuits could potentially be holding solutions to increasing the speed of computing beyond silicon computing.















I did not understand a single thing… Time to up my game & get up to speed… But but DNA computers?
Code is code… Learn the language and compile.
Pretty easy really. Just get together but loads of hydrogen and wait.
Eventually stars and planets form some are habitable and life evolves.
Not long later the first graduate students appear and might solve your problem for you!
Now that’s an evolutionary algorithm!
That’s already been done.
The answer is 42
https://www.popularmechanics.com/science/a30455677/chemical-turing-machine/
Only Intel is struggling at 10nm. The rest of the industry uses TSMC, they’re at 7nm, and not struggling. Iut will be interesting to see where Samsung’s new 250 Billion Dollar plant can get to, But I think its safe to say foundries are beyond company scale now and are nation state projects. TSM has the right model, be a foundry, and make it available to everyone. That’s the only way to amortize those kinds of costs.
But can it run Doom?
You don’t want to know!
You certainly don’t want the scientists trying that!
Is this really calculating a value? it looks like they pre-load the sequence with values that once acted on my the enzymes can only produce the desired result. It would seem the calculation is really external. Meaning you needed to know the number in advance and load that information into the pairs. The inclusion of a truth-table is very fitting. Yes is can be programmed and indicative, but the truth-table is not a computer.
That’s essentially what all digital logic looks like. It’s certainly what the logic in the processor of whatever device you used to send that comment looks like.
How many MHz does it run at? 0.00000004? I don’t think our current models of computing adapt well to this brave new world.
The clock speed isn’t all that great, but half a liter of this stuff could give you a core count in the hextillions. :-)
Yeah not fast, but massively parallel processes are possible even in a single cell.
http://book.bionumbers.org/what-is-faster-transcription-or-translation/
I’d like to see them use the artificial base pairs in Hachimoji dna in a dna computer, it would mean less possibility of interference with natural biological systems in both directions.
How can dna be 10 times smaller than 10nm?!
A silicon atom is 6nm. How can dna be smaller than Si atom?!
Are you sure you’re not thinking of the spacing of silicon atoms in an ionic lattice?
Is it wrong to think this? (I don’t know.) Why?
Hmm, just double checked and Si is .11nm. Still DNA has thousands of atoms doesn’t it? How can it fit in 1nm?
The DNA molecule is about 2nm WIDE. However, they are often meters long.
That might sound like an unfair comparison, but not really. The “process size” quoted is the transistor’s “length”, or the minimum length of the gate over the channel. The actual useful transistor size can vary wildly depending on how it is used, but is going to be significantly larger.
Still seems like an apples-to-oranges comparison.