In the blue corner, we have the VENUS FLYTRAP! In the red corner, we have the underdog of the century, AN ENTIRE PARTICLE ACCELERATOR. Yes, you read that right. When you have a particle accelerator, it’s only second nature to throw anything you can into it. That’s why [Electron Impressions] put a poor fly-eating trap into their accelerator.
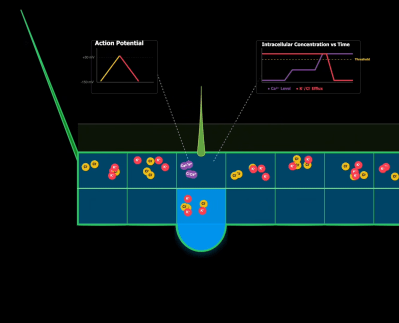
The match-up isn’t quite as arbitrary as it might seem at first. The flytrap’s main mechanism of trapping and digesting insects relies heavily on intracellular ion movement. Many cells along the inside of the trap have hair-activated calcium channels that respond to a fly landing on its surface. This ion movement then creates an action potential, which propagates along the entire surface, triggering closing. As the potential moves across different cells, other ions leave and create osmotic pressure. This pressure is what creates the mechanical movement.
Of course, this makes it no surprise when the plant finds itself under the ionizing radiation that every single head closes at once. While this is a cool demonstration, there is a slight side effect of killing every single cell by ripping apart the trap’s DNA.
Well, who would have guessed that the underdog accelerator would have won… Anyways, the DNA being ripped apart is far from ideal for repeatability. If you want to learn more about genetic features that SHOULD be repeated, then make sure to check out the development of open-source insulin!
Continue reading “Venus Flytrap Takes Ride Through A Particle Accelerator”

















