[Bruce Land] is a professor at Cornell University who was looking for a way to quickly solve chemical kinetic systems. He had used MATLAB but longed for a faster method. His upgrade achieves a 100 times speed increase by using an FPGA as a parallel stochastic solver.
It works by generating 100 pseudo-random 16-bit numbers using an Altera DE2 board. This is done once per cycle at 50 MHz so we’re talking about a lot of random numbers. They are run through the solver algorithms and used to compute each reaction cycle. On a 3.8 GHz P4 process running the MATLAB version one of these cycles would take about 1000 seconds, so the speed improvement can immediately be felt. It’s wonderful to have this new tool. It does make us wonder what could be done with GPU processing that we’ve seen for password cracking or bit coin mining. Much like FPGAs a GPU is prefect for running a large number of parallel operations.

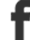













1.1 seconds on a FPGA, 1000 seconds on the P4 (sorry if that was obvious to everyone else)
http://forum.bitcoin.org/index.php?topic=9047.0
bitcoin mining with an FPGA
http://nsa.unaligned.org/
sha1/md5 hashing on an FPGA for password cracking
Sometimes these writeups are almost unintelligible. I know it’s kind of cliché to decry the quality of HaD posts, but come on.
@Ioans: agreed. There are very few write-ups that don’t have issues thr make them difficult to read. Glad I’m not the only one to notice it.
Otherwise, very interesting use for FPGAs.
@spadusa: HAD don’t care – see this thread http://forums.hackaday.com/viewtopic.php?f=2&t=858
Very interesting!
Bruce Land is a legend. He is the one behind almost all of Cornell projects that get posted here: http://people.ece.cornell.edu/land/courses/ece4760/FinalProjects/
It’s actually rather difficult to generate the quantity of random numbers needed to make use of a GPU for this sort of task.
This is very interesting and I’ll have to check it over very carefully.
The tough thing about chemical simulations is the number of iterations required and the magnitude of the numbers being multiplied: 15 orders of magnitude change in concentration wouldn’t be unusual, and if you’re simulating millions of reactions, errors stack up fast.
This was my first eye-opener for IEEE 754.