Improvements in methodology have dramatically dropped the cost of DNA sequencing in the last decade. In 2007, it cost around $10 million dollars to sequence a single genome. Today, there are services which will do it for as little as $1,000. That’s not to bad if you just want to examine your own DNA, but prohibitively expensive if you’re looking to experiment with DNA in the home lab. You can buy your own desktop sequencer and cut out the middleman, but they cost in the neighborhood of $50,000. A bit outside of the experimenter’s budget unless you’re Tony Stark.
 But thanks to the incredible work of [Alexander Sokolov], the intrepid hacker may one day be able to put a DNA sequencer in their lab for the cost of a decent oscilloscope. The breakthrough came as the result of those two classic hacker pastimes: reverse engineering and dumpster diving. He realized that the heavy lifting in a desktop genome sequencer was being done in a sensor matrix that the manufacturer considers disposable. After finding a source of trashed sensors to experiment with, he was able to figure out not only how to read them, but revitalize them so he could introduce a new sample.
But thanks to the incredible work of [Alexander Sokolov], the intrepid hacker may one day be able to put a DNA sequencer in their lab for the cost of a decent oscilloscope. The breakthrough came as the result of those two classic hacker pastimes: reverse engineering and dumpster diving. He realized that the heavy lifting in a desktop genome sequencer was being done in a sensor matrix that the manufacturer considers disposable. After finding a source of trashed sensors to experiment with, he was able to figure out not only how to read them, but revitalize them so he could introduce a new sample.
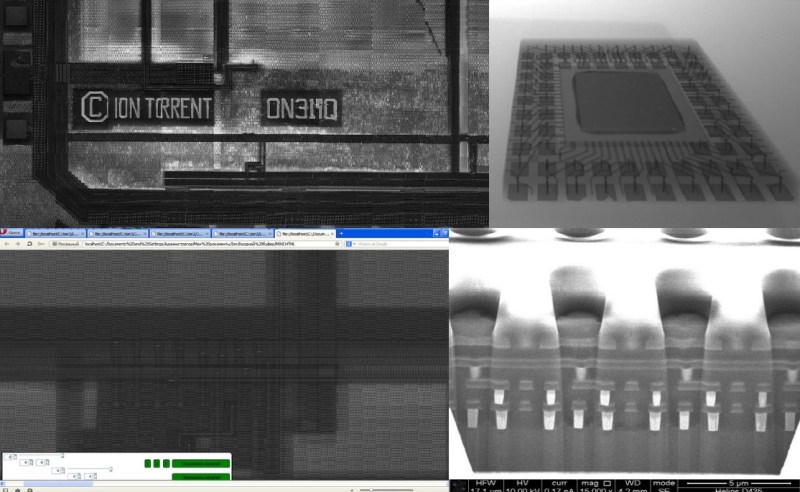
To start with, [Alexander] had to figure out how these “disposable” sensors worked. He knew they were similar in principle to a digital camera’s CCD sensor; but rather than having cells which respond to light, they read changes in pH level. The chip contains 10 million of these pH cells, and each one needs to be read individually hundreds of times to capture the entire DNA sequence.
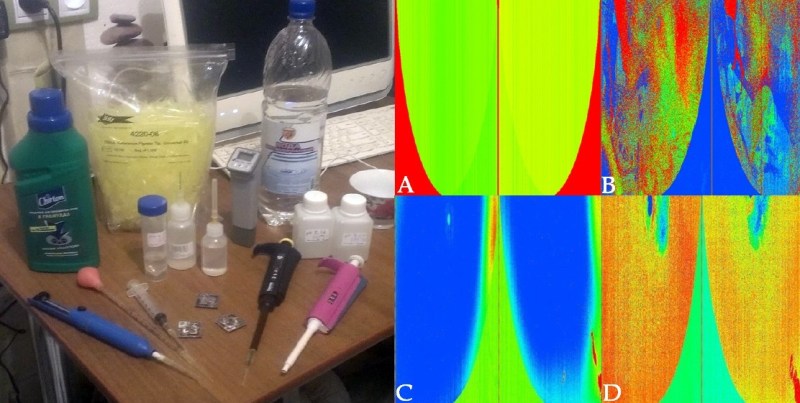 Enlisting the help of some friends who had experience reverse engineering silicon, and armed with an X-Ray machine and suitable optical microscope, he eventually figured out how the sensor matrix worked electrically. He then designed a board that reads the sensor and dumps the “picture” of the DNA sample to his computer over serial.
Enlisting the help of some friends who had experience reverse engineering silicon, and armed with an X-Ray machine and suitable optical microscope, he eventually figured out how the sensor matrix worked electrically. He then designed a board that reads the sensor and dumps the “picture” of the DNA sample to his computer over serial.
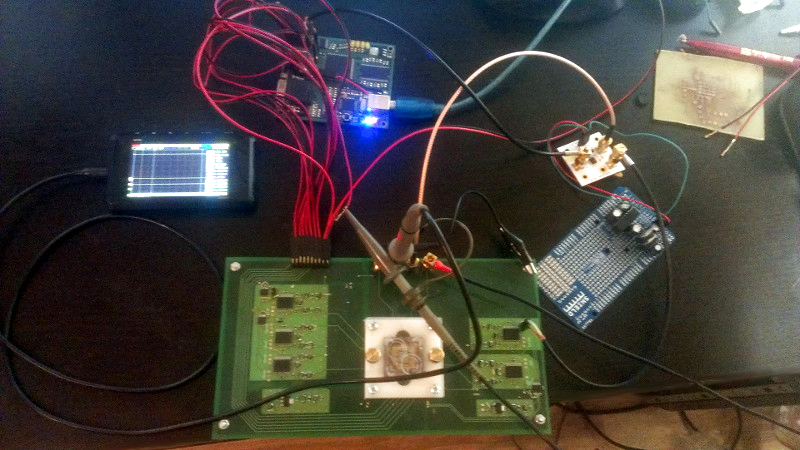
Once he could reliably read the sensor, the next phase of the project was finding a way to wash the old sample out so it could be reloaded. [Alexander] tried different methods, and after several wash and read cycles, he nailed down the process of rejuvenating the sensor so its performance essentially matches that of a new one. He’s currently working on the next generation of his reader hardware, and we’re very interested to see where the project goes.
This isn’t the first piece of DIY DNA hardware we’ve seen here at Hackaday, and it certainly won’t be the last. Like it or not, hackers are officially fiddling with genomes.














Not a hack. This goes way beyond a hack. This is magic. Very impressive.
“Improvements in methodology have dramatically dropped the cost of DNA sequencing in the last decade. In 2007, it cost around $10 million dollars to sequence a single genome. Today, there are services which will do it for as little as $1,000. ”
Only took eleven years though.
“That’s not to bad if you just want to examine your own DNA, but prohibitively expensive if you’re looking to experiment with DNA in the home lab.”
Cheaper than DIY trans-humanism.
Just gotta nail 1 pot dA-tail and ligation. Apparently its just a few polymerases ;)
Also anybody know what fragmentase is?
:P
“Also anybody know what fragmentase is?”
It’s when the cops taze a perp to pieces?
B^)
The man starts his article by stating he’s an expert in “quantum cryptography”… He should say “quantum teleportation” instead. It rocks better.
Great article, I wonder how much those sensors are that the manufacturer says you have to throw away :-P
“Improvements in methodology have dramatically dropped the cost of DNA sequencing in the last decade. In 2007, it cost around $10 million dollars to sequence a single genome. Today, there are services which will do it for as little as $1,000. ”
The “$” means “Dollars”, so either leave it off and say “10 million dollars” or drop the “dollars” and leave the $ (And most style guides say either is correct, but to be consistent)
“That’s not to bad if you just want to examine your own DNA, but prohibitively expensive if you’re looking to experiment with DNA in the home lab.”
too bad, not to bad (unless there’s a place called “bad” and you’re going to bad :-P
no STEM cell ;)
fuuuu…. *detected* …and the joke is burnt :(
*detected*
Well, I am thinking that the mfgr wants the trays to be junked because of the possibility of ANY contamination of results by a DNA fragment from a previous test.
It the cleansing process can guarantee that no fragments remain, okay.
s/It/If
Cleanliness is vital. Wouldn’t want the molecular equivalent of what happened to Archibald Buttle. So for criminal and paternity tests they’re always (should always!) going to use a new sensor.
In theory you can pair different samples with different DNA “barcodes” so you know whether the read is from the new sample or an old sample, but the manufacturer tends to want you to buy new things rather than reuse old things. Oxford Nanopore actually sell a kit for washing their flowcells: https://store.nanoporetech.com/flow-cell-wash-kit-r9.html
Truly awesome. I didn’t see any sample DNA sequence readouts in the discussion though. Does he know the accuracy of the reads, or is it still early days?
A desktop sequencer from oxford nanopore is $1,000, not $50k. And no hacking required
How much do consumables for it cost per read?
Following Fay’s link suggests that can do 200kb reads. The chip this guy is hacking does millions of (hundreds or thousands of base pair per cell) reads in parallel. I’m not clear on how each cell is filled. That sounds like massive automation needed to me.
You’re confusing the number of reads with the length of the reads. The Nanopore can theoretically produce reads 200Kbp long, whereas the Ion Torrent is something like 400bp long. The Nanopore can apparently keep producing reads for as long as there is DNA on the right side of the flow cell, whereas the Ion torrent will be fixed at a few hundred million reads output depending on how it is run.
Someone where I work said that their Nanopore run gave reads which were 10Kbp long, which is far short of the advertised value, but is still far longer than any other machine produces. The error rate is comparatively high, however.
Err $1,000 commercial Nanopore sequencer anyone? https://nanoporetech.com/products/minion