Money, status, or even survival – there’s no shortage of incentives for faking results in the scientific community. What can we do to prevent it, or at least make it noticeable? One possible solution is cryptographic signing of measurement results.
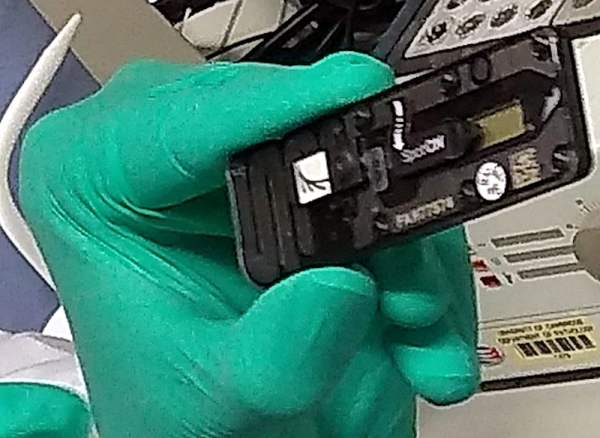
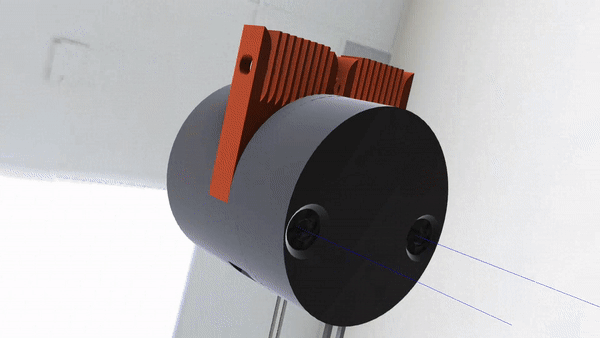
Here’s a proof-of-concept from [Clement Heyd] and [Arbion Halili]. They took a ThermoFisher Scientific 7500 Fast PCR (Polymerase Chain Reaction) machine, isolated its daughter-software, and confined it into a pipeline that automatically signs each result with help of a HSM (Hardware Security Module).
A many machines do, this one has to be paired to a PC, running bespoke software. This one’s running Windows XP, at least! The software got shoved into a heavily isolated virtual machine running XP, protected by TEE (Trusted Execution Environment). The software’s output is now piped into a data diode virtual serial port out of the VM, immediately signed with the HSM, and signed data is accessible through a read-only interface. Want to verify the results’ authenticity? Check them against the system’s public key, and you’re golden – in theory.
This design is just a part of the puzzle, given a typical chain of custody for samples in medical research, but it’s a solid start – and it happens to help make the Windows XP setup more resilient, too.
Wondering what PCR testing is good for? Tons of things all over the medical field, for instance, we’ve talked about PCR in a fair bit of detail in this article about COVID-19 testing. We’ve also covered a number of hacker-built PCR and PCR-enabling machines, from deceivingly simple to reasonably complex!